)

Antibiotic use kills susceptible bacteria, but resistant strains survive - meaning that infections can get harder to treat.
For the past three generations, antibiotics have enabled a whole realm of possibilities within medicine.
Things that we now consider routine wouldn’t even have been possible 100 years ago, owing to the infection risk, even if they’d been technically possible. Lung transplants, bone marrow transplants, intensive care medicine, gut surgery more complex than having an appendix removed…all of these would have carried an overwhelming risk of infection to the patient, with a near-certainty of death.
In short, our ability to treat infection allows medicine to do things that were previously considered to be unthinkable.
However, with the use of antibiotics comes a new problem. Although we can treat patients after surgery or illness, this in turn has steadily increased the pool of vulnerable patients. These folk develop more infections and, as antibiotics are used to treat them, antibiotic-resistant bacteria are selected.

Why are bacteria becoming more resistant to antibiotics?
Bacteria breed quickly and exchange genetic information, which means that they have many different routes to develop resistance to antibiotics.
Over time, many types of bacteria have become more resistant to antibiotics. In the early 1960s ampicillin was reliable against E. coli, which is the commonest type of bacteria in urinary and bloodstream infections, but nowadays 60% of E. coli are ampicillin-resistant and 10-12% are resistant to the cephalosporins developed to replace ampicillin.
Antibiotic resistance has become a major international public health issue, with fears that bacteria are becoming resistant faster than pharma is developing new antibiotics. There are two particular areas of risk where we might run out of antibiotic treatments in the foreseeable future.
The first is in specialist settings, such as intensive case and with immune-suppressed patients. These patients are vulnerable to many opportunist bacteria that are very adept at gaining resistance. The second area of risk comes in several classical infectious diseases such as TB, typhoid and gonorrhoea, where accumulating risk means that we’re already down to very few treatment options.
Even for the most resistant bacteria there’s usually still some antibiotic that’s still active. However, current methods mean that it takes two days to grow the patient’s bacteria in a laboratory and discover which antibiotics kill them. While this is happening, the patient is treated blind using a broad-spectrum antibiotic to cover all likely bacteria in the type of infection. There are two flaws with this approach – first, there’s the risk that first empirical antibiotic turns out not to be active. This risk is greatest in countries where resistance is most prevalent. Second, there’s the hazard that the patient is being over-treated when, in fact, they have a susceptible pathogen. The result is an unnecessary selection for resistance among the patient’s gut bacteria, which may be a source of future infections.
“Resistance is the greatest demonstration of Darwin’s survival of the fittest – we’ve been very good at killing susceptible bacteria, but resistant ones survive and infect further patients.”
-Dr David Livermore
There are four key ways to help minimise the risk and impact of antibiotic resistance:
1. Good Infection Control
Stopping people from getting infections in the first place means that we use antibiotics less, reducing the selection of new resistance. Prevention is multi-faceted and includes good hand washing in hospitals, using condoms to stop the spread of sexually-transmitted diseases, and vaccines.
The more we can do to reduce infection, the fewer patients will require antibiotics and the less selection of resistance.
2. Development of New Antibiotics
In the past, the development of new antibiotics was sufficient to keep ahead of resistance. However, it’s become increasingly difficult - new antibiotics are technically hard to discover and there’s a misalignment between what the regulators require to grant a license and what clinicians seek. What’s more, antibiotics aren’t a very attractive area for development commercially: they are taken for a week or two whereas heart medicines are for life. With these factors in mind, we need to turn to new means to fight infections.
3. Stewardship
The principle of ‘stewardship’ is that, in order to maintain the effectiveness of antibiotics, we need to balance the needs of patients with the need to conserve our stock of antibiotics, avoiding unnecessary use and selection for resistance.
Put simply it’s the laudable aim of ‘Right drug, right dose, right duration’ for patients who need antibiotics and avoiding use in those who don’t need them.
The problem, again, is that the clinician doesn’t know the type of bacteria, nor their resistances until 2 or 3 days after he or she starts to treat the infection…. And this drives the precautionary use of broad-spectrum agents.
4. Rapid diagnosis
By shortening the time it takes to identify pathogens and their antibiotic resistance, patients will be given more effective and appropriate treatment. As well as helping the patient, this also has a wider potential benefit to society as antibiotic wastage is reduced.
This is the aspect of tackling antibiotic resistance that the INHALE project is focusing upon.


INHALE
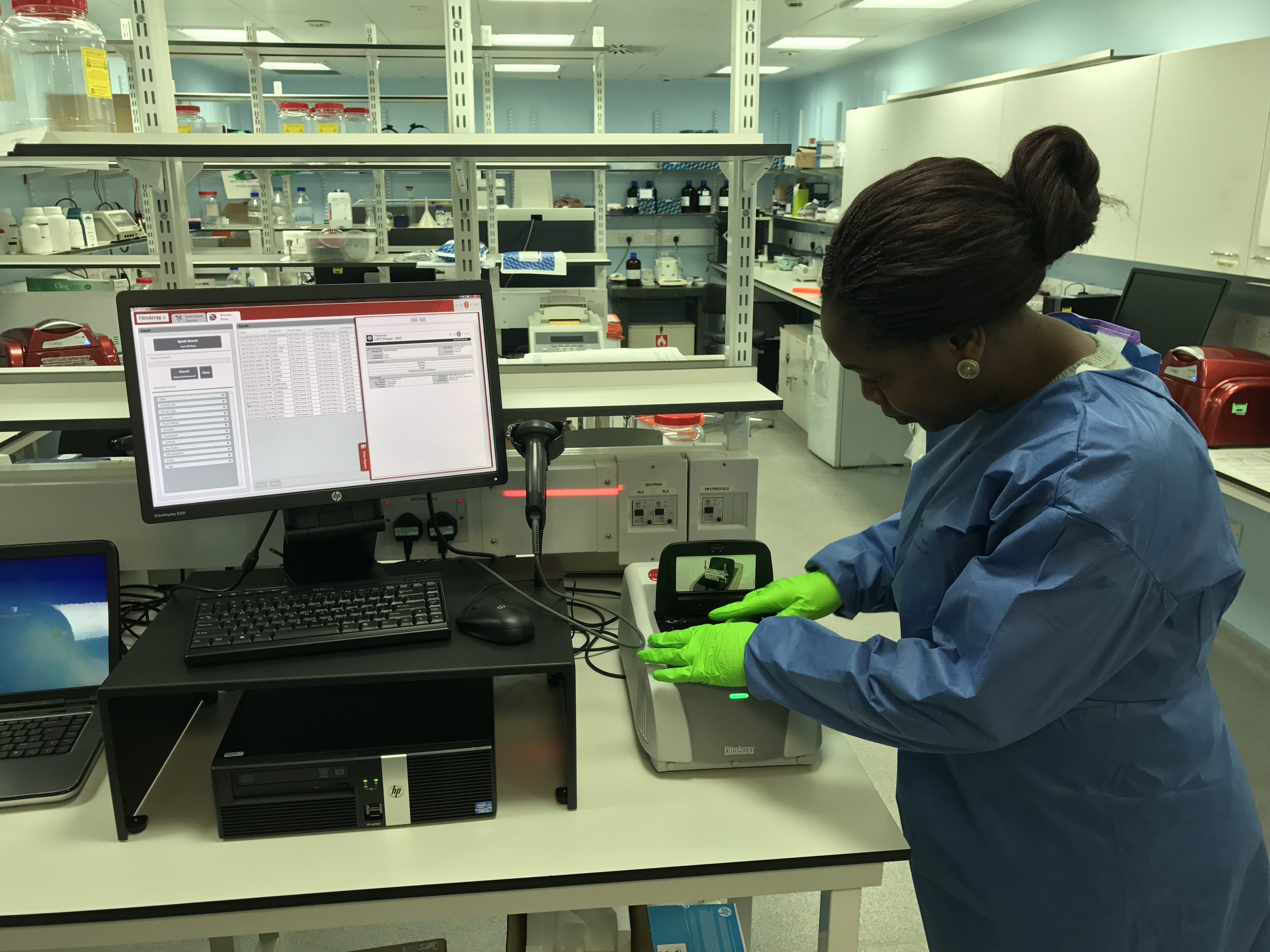
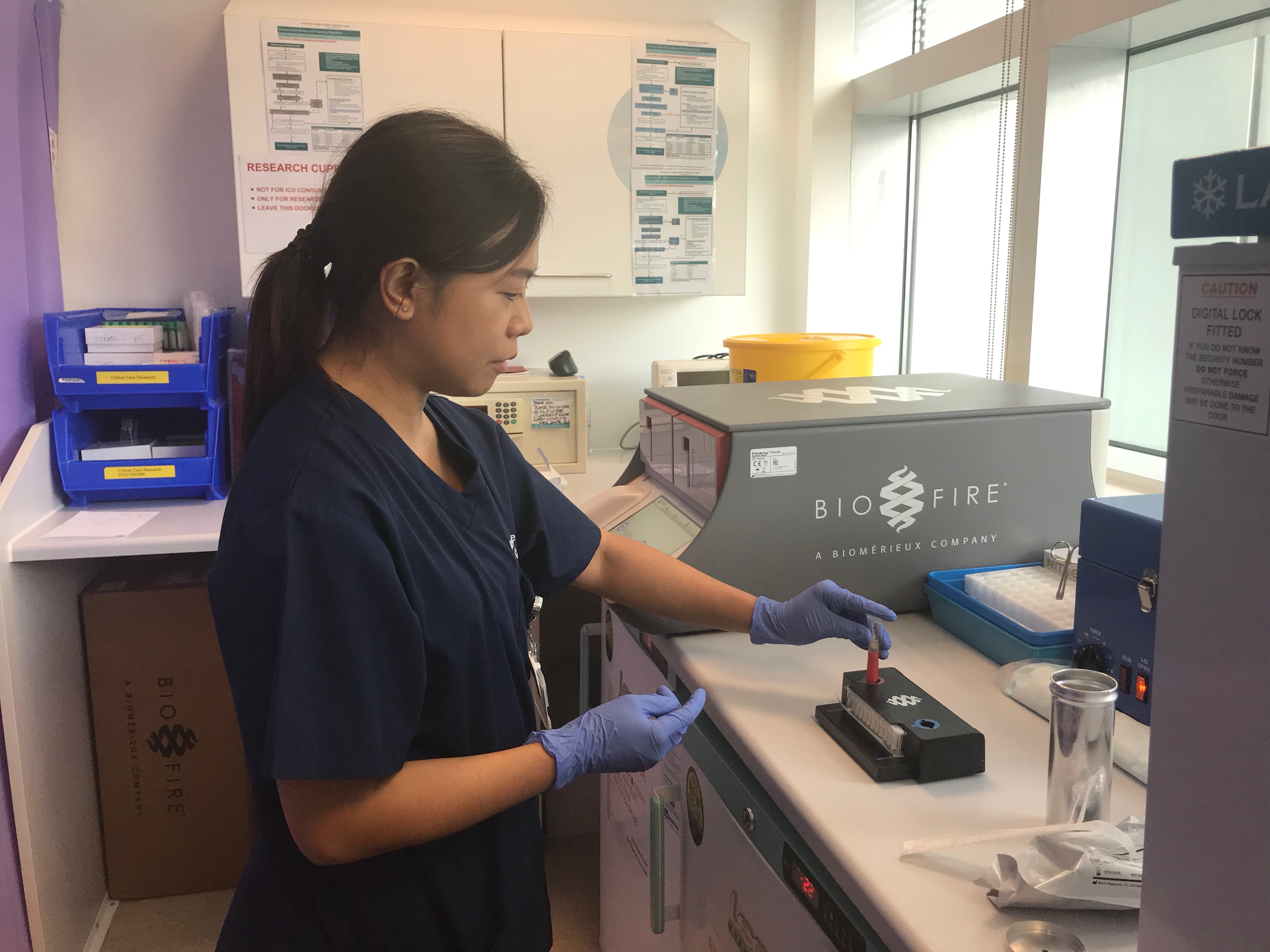
To help shorten the delay in identifying pathogens and their antibiotic resistance, INHALE, funded by a Programme Grant from the UK National Institutes for Health Research, is exploring the potential of a rapid molecular diagnostic – the BioFire FilmArray Pneumonia Panel - in hospital-acquired and ventilator-associated pneumonia (HAP and VAP). These are the commonest infections developed by patients in intensive care units (ICUs).
The FilmArray diagnostic was chosen as the preferred tool in a first phase of INHALE. This took place from 2016 – 2018 and involved taking respiratory secretions from pneumonia patients and putting them through several different molecular systems. The researchers then picked the best system, both for agreement with conventional microbiology and ease of use.
The trial is running across 12 UK hospitals, 11 NHS and one private. Consented HAP or VAP patients in the ICU are randomised either to be treated using standard best practice (where they’ll receive a broad spectrum antibiotic until the lab can identify the pathogen and prescribe a more tailored treatment, after 48-72 hours), or to have their treatment guided by the FilmArray molecular test, which gives a result in a little over an hour.
In principle, and if clinicians act on the result, this allows patients to be given a narrow-spectrum antibiotic almost from the outset of treatment.
Professor David Livermore is a Co-Chief Investigator on the project.
He’s enthusiastic about the trial and its possibilities;
“We’ve currently recruited 92 patients and the recruitment rate means that we’re on course to reach our target of 466. This is a hugely exciting project, as the test we’re looking at can be done by the nurse in the ICU – it doesn’t have to go to a lab to be processed, although results must still be discussed with the microbiology department to ensure best stewardship.”
The INHALE project will run until 2021, with two main outcome measures:
- To ensure patient outcomes are at least as good as with present standard-of-care
- To improve antibiotic stewardship – this technology should lead to less use of broad spectrum antibiotics, and increased use of narrow spectrum antibiotics targeted precisely against the patient’s pathogen.
The INHALE project aims to improve antibiotic use in HAP and VAP. The current system of using broad-spectrum antibiotics in all cases isn’t ideal because of the risk of resistant bacteria being selected in the guts of patients who don’t have unusually resistant bacteria causing the infection in their lungs. Rapid recognition of pathogens and their resistances provides the most attractive route to tackle antibiotic resistance, aiming to tighten use stewardship whilst maintaining or improving patient outcomes.
Future direction
Although the INHALE project is still ongoing, molecular testing of bacteria, without the delays of conventional lab culture, could potentially have far reaching consequences beyond HAP and VAP patients. The same principles and methodologies may be applicable to other infections (UTIs for example), with results used to develop prescribing strategies and 'rules' for best antibiotic use.
INHALE is managed by a multidisciplinary team of researchers and clinicians from two universities (UCL and UEA) and currently 12 ICUs are recruiting patients.
Further information:
https://www.ucl.ac.uk/inhale-project