Given a set of polyploids coupled with their corresponding ploidy levels, i.e., the number of complete chromosome sets found in the cells of an organism, SPRINT computes in most cases the minimum number of hybrid vertices in a phylogenetic network which can explain the ploidy levels. In addition, a figure of a phylogenetic network which exhibits this minimum number is displayed within the SPRINT GUI, with the option to export this network in DOT format to the desired analytical and visualisation tool of choice (e.g. Cytoscape).
Availability
SPRINT is freely available and may be accessed online at https://github.com/lmaher1/SPRINT or downloaded as a zip file which contains:
The Python source code, SPRINT.py.
An alternative version of SPRINT which does not require pygraphviz (and thus Homebrew) to be installed, SPRINT-nxspring.py.
A requirements file containing four pieces of software able to be installed using a pip install.
The SPRINT Manual (PDF, 682 KB) which details the installation and running of SPRINT.
A tests folder containing four .txt files. One of which is the dataset of Viola species used in Huber and Maher (2022) and also in the SPRINT manual. The remaining tests are fictional examples used to ensure the program is running as intended.
Install instructions (txt)
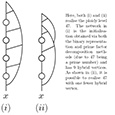
non optimal initialisation
Support
In case of any questions, email Katharina T. Huber.
Related papers
Huber, K.T., Maher, L.J. The hybrid number of a ploidy profile. J. Math. Biol. 85, 30 (2022). https://doi.org/10.1007/s00285-022-01792-6
Research Team

)